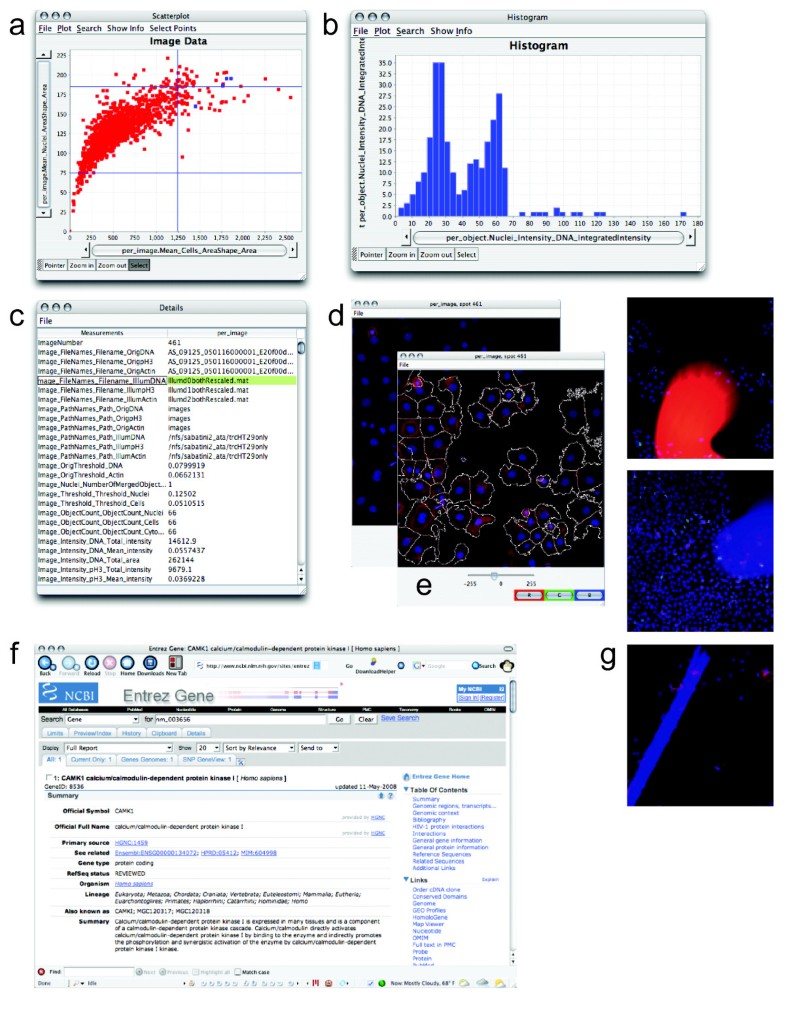
You should also view the predicted segmentation, and annotate pixels that are not currently segmented properly.Ĭontinue until it seems that additional labels do not change the results, or a subset of the pixels begin “flipping” between CHO cell and background. Areas of high uncertainty will be aqua-blue, so annotating those areas will be most beneficial to training the program which pixels belong to which class. Toggling the Segmentation and Uncertainty views provides real-time feedback that can guide the labeling process. This outcome is promising considering this classification was determined by 1 feature and 1 pixel each for the CHO and background labels.Īdding more labels, one pixel at a time, we continue to refine the segmentation. Activating Live Update reveals the segmentation looks similar to the results from thresholding. In the next image, the annotation color of the CHO cell is yellow and the annotation color of the background is green. Using a brush size of 1, we click a single pixel from each class: one within a single CHO cell and the other in the surrounding background. The image below shows how closely we must view individual cells before the pixels of the image become clear. To label one pixel at a time, we’ll need to zoom in far enough to resolve the individual pixels in the image. too much information about any given area can cause classification to do poorly in other slightly-dissimilar areas. We recommend creating labels for each class one pixel at a time, rather than by making scribbles, to minimize the chance of over-fitting, i.e. We will begin here by labeling pixels for two classes: a background class and a CHO cell class. Zoom-in far enough that the grid is no longer visible.The keyboard shortcut Ctrl-D will show the grid Ilastik uses to partition the image for processing.Some shortcuts that may prove useful are: Open ilastik, load an image, and seek out a cell that looks representative of the population. This annotation creates what is referred to in machine learning as a training set. ilastik provides a user interface for labeling, tagging, and identifying the objects of interest within an image. The machine-learning implemented by ilastik requires user annotation about what is background and what is a CHO cell before it can automatically make this determination across a set of images. These same patterns can be detected with the machine-learning algorithms within ilastik. The CHO cells within the DIC image are obvious to the human eye, because we can discern that each cell is defined by a characteristic combination of light and dark patterns. Ilastik employs pixel-based classification and complements CellProfiler. But let’s pretend for a moment we didn’t have that option… Now, note that there is a module, EnhanceOrSuppressFeatures, that is specifically capable of transforming DIC images into something that is readily segmented. 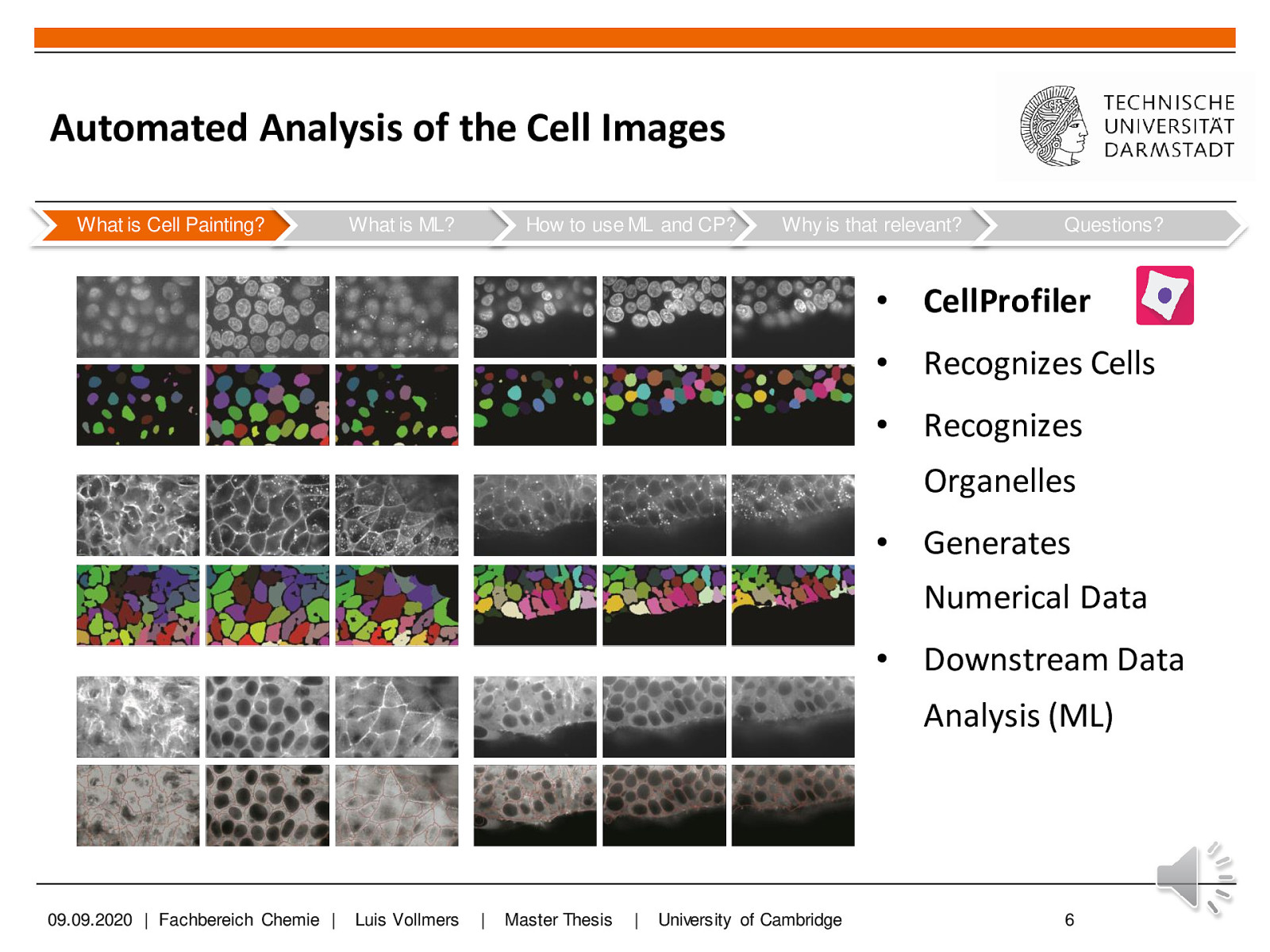
While these fragments could be useful for simple cell counting, most metrics of morphology will be inaccurate. Thresholding renders the CHO cells into moon-like crescents. There is therefore no true foreground for these cells based solely upon an intensity histogram. The goal will be to identify individual Chinese Hamster Ovary (CHO) cells and the regions they occupy.Ī straightforward thresholding of this image yields poor results, because the cells have almost the same pixel intensity values (and sometimes even darker!) as the background. Now, let’s take a look at how ilastik can be used together with CellProfiler! DIC conundrumĬonsider segmenting DIC images, such as those within the imageset BBBC030. ilastik is an open-source tool built for pixel-based classification, and, when combined with CellProfiler, the range of biology that can be quantified from images is greatly expanded beyond monocultures of monolayers to include increased complexity such as tissues, organoids, or co-cultures. Thankfully, machine learning, particularly pixel-based classification has yielded powerful techniques that can often solve these challenging cases. For instance, when the biologically relevant objects are defined more by texture and context than raw intensity many classical image processing techiques can be foiled DIC images of cells are a common biological example. However, despite the many problems CellProfiler can readily solve, certain types of images are particularly challenging. Peruse the documentation on the IdentifyPrimaryObjects module, for example, to get a sense of these, e.g., thresholding, declumping, and watershed. CellProfiler is capable of accurate and reliable segmentation of cells by utilizing a broad collection of classical image processing methods.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
